New DNA Analysis Says You've Got More Pre-Human Ancestors
This week a research paper published in PLOS Genetics revealed a newly developed algorithm for analyzing genomes. Researchers used their new system on the genomes of two Neanderthals, two humans of African descent, and a Denisovan. Approximately 1% of the genome of the Denisovan came from an "unsequenced, but highly diverged, archaic hominin ancestor."
This previously unknown "super-archaic hominin" crossed over after chimps, but before Hominin split into Archaic and what became Human. It's suggested by this research that "about 15% of these "super archaic regions-comprising at least about 4Mb-were, in turn, introgressed into modern humans and continue to exist in the genomes of people alive today."
As such, we've got ancestors we did not even know existed until this research was published. The research notes that they've found "no convincing evidence that selection acted against any of these introgressed regions." In other words, those ancient beings coupled with these previously unknown beings and popped out babies that then popped out more babies, and so on until today, where you and I almost certainly have ancient, unknown being DNA!
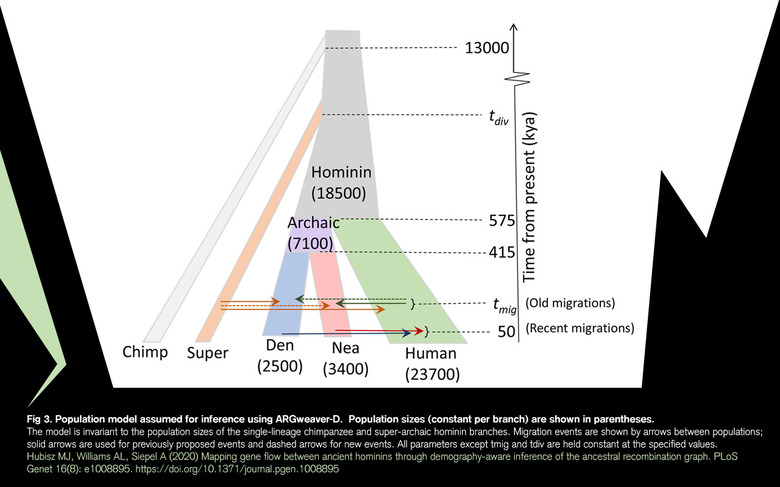
"What I think is exciting about this work is that it demonstrates what you can learn about deep human history by jointly reconstructing the full evolutionary history of a collection of sequences from both modern humans and archaic hominins," said Adam Siepel, one of three authors of the new research
"This new algorithm that Melissa has developed, ARGweaver-D, is able to reach back further in time than any other computational method I've seen," said Siepel via Eurekalert. "It seems to be especially powerful for detecting ancient introgression."
To learn more about this subject, head over to the paper Mapping gene flow between ancient hominins through demography-aware inference of the ancestral recombination graph. This paper was published with code DOI: 10.1371/journal.pgen.1008895 on August 6, 2020. Authors of this paper include Melissa J. Hubisz, Amy L Williams, and Adam Siepel.
